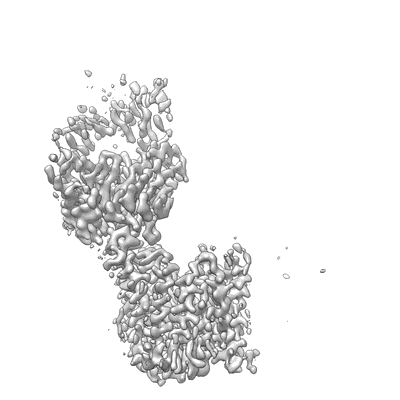
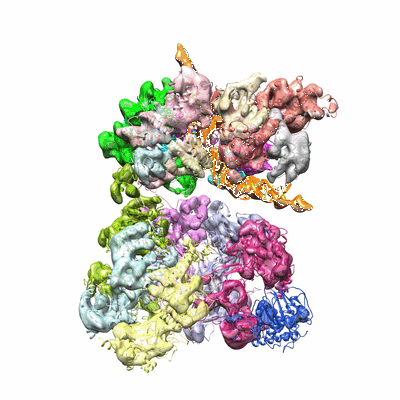
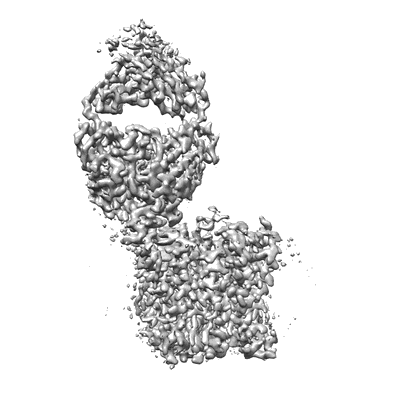CryoEM structure of the symmetric Pho90 dimer from yeast without substrates.
Resolution: 2.29 Å
EM Method: Single-particle
Fitted PDBs: 8r33
Q-score: 0.676
Schneider S, Kuhlbrandt W, Yildiz O
Structure (2024) [ Pubmed: 38688287 DOI: doi:10.1016/j.str.2024.04.005 ]
- Pho90 isoform 1 (97 kDa, Protein from Saccharomyces cerevisiae)
- Water (18 Da, Ligand)
- 1,2-diacyl-glycerol-3-sn-phosphate (704 Da, Ligand)
- Dimeric scpho90 (Complex from Saccharomyces cerevisiae)
CryoEM structure of the symmetric Pho90 dimer from yeast with substrates.
Resolution: 2.62 Å
EM Method: Single-particle
Fitted PDBs: 8r34
Q-score: 0.624
Schneider S, Kuhlbrandt W, Yildiz O
Structure (2024) [ Pubmed: 38688287 DOI: doi:10.1016/j.str.2024.04.005 ]
- Phosphate ion (94 Da, Ligand)
- Low-affinity phosphate transporter pho90 (97 kDa, Protein from Saccharomyces cerevisiae)
- Dimeric scpho90 (Complex from Saccharomyces cerevisiae)
- Sodium ion (22 Da, Ligand)
- 1,2-diacyl-glycerol-3-sn-phosphate (704 Da, Ligand)
- Water (18 Da, Ligand)
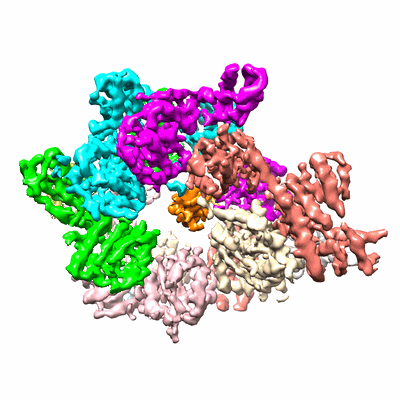
CryoEM structure of the asymmetric Pho90 dimer from yeast without substrates.
Resolution: 3.12 Å
EM Method: Single-particle
Fitted PDBs: 8r35
Q-score: 0.529
Schneider S, Kuhlbrandt W, Yildiz O
Structure (2024) [ Pubmed: 38688287 DOI: doi:10.1016/j.str.2024.04.005 ]
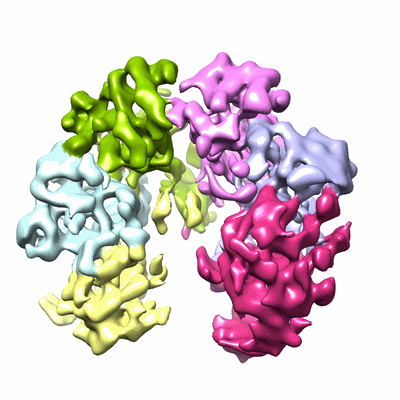
Structure of the human 20S U5 snRNP core
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 8q91
Q-score: 0.417
Schneider S, Brandina I, Peter D, Lagad S, Fraudeau A, Portell-Montserrat J, Tholen J, Zhao J, Galej WP
Nat Struct Mol Biol (2024) [ Pubmed: 38467877 DOI: doi:10.1038/s41594-024-01250-5 ]
- Small nuclear ribonucleoprotein sm d1 (13 kDa, Protein from Homo sapiens)
- Probable atp-dependent rna helicase ddx23 (95 kDa, Protein from Homo sapiens)
- U5 snrna (37 kDa, RNA from Homo sapiens)
- U5 small nuclear ribonucleoprotein 200 kda helicase (244 kDa, Protein from Homo sapiens)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Small nuclear ribonucleoprotein g (8 kDa, Protein from Homo sapiens)
- Pre-mrna-processing-splicing factor 8 (273 kDa, Protein from Homo sapiens)
- 116 kda u5 small nuclear ribonucleoprotein component (109 kDa, Protein from Homo sapiens)
- Small nuclear ribonucleoprotein f (9 kDa, Protein from Homo sapiens)
- Small nuclear ribonucleoprotein e (10 kDa, Protein from Homo sapiens)
- Small nuclear ribonucleoprotein-associated proteins b and b' (24 kDa, Protein from Homo sapiens)
- Cd2 antigen cytoplasmic tail-binding protein 2 (37 kDa, Protein from Homo sapiens)
- Small nuclear ribonucleoprotein sm d2 (13 kDa, Protein from Homo sapiens)
- 20s u5 snrnp complex purified from hek293f cell line (1 MDa, Complex from Homo sapiens)
- Pre-mrna-processing factor 6 (107 kDa, Protein from Homo sapiens)
- Small nuclear ribonucleoprotein sm d3 (13 kDa, Protein from Homo sapiens)
Structure of the human 20S U5 snRNP
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 8rc0
Q-score: 0.309
Schneider S, Brandina I, Peter D, Lagad S, Fraudeau A, Portell-Montserrat J, Tholen J, Zhao J, Galej WP
Nat Struct Mol Biol (2024) [ Pubmed: 38467877 DOI: doi:10.1038/s41594-024-01250-5 ]
- Small nuclear ribonucleoprotein sm d1 (13 kDa, Protein from Homo sapiens)
- Probable atp-dependent rna helicase ddx23 (95 kDa, Protein from Homo sapiens)
- U5 snrna (37 kDa, RNA from Homo sapiens)
- U5 small nuclear ribonucleoprotein 200 kda helicase (244 kDa, Protein from Homo sapiens)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Small nuclear ribonucleoprotein g (8 kDa, Protein from Homo sapiens)
- Pre-mrna-processing-splicing factor 8 (273 kDa, Protein from Homo sapiens)
- U5 small nuclear ribonucleoprotein 40 kda protein (39 kDa, Protein from Homo sapiens)
- 116 kda u5 small nuclear ribonucleoprotein component (109 kDa, Protein from Homo sapiens)
- Small nuclear ribonucleoprotein f (9 kDa, Protein from Homo sapiens)
- Small nuclear ribonucleoprotein e (10 kDa, Protein from Homo sapiens)
- Small nuclear ribonucleoprotein-associated proteins b and b' (24 kDa, Protein from Homo sapiens)
- Cd2 antigen cytoplasmic tail-binding protein 2 (37 kDa, Protein from Homo sapiens)
- Small nuclear ribonucleoprotein sm d2 (13 kDa, Protein from Homo sapiens)
- 20s u5 snrnp complex purified from hek293f cell line (1 MDa, Complex from Homo sapiens)
- Pre-mrna-processing factor 6 (107 kDa, Protein from Homo sapiens)
- Small nuclear ribonucleoprotein sm d3 (13 kDa, Protein from Homo sapiens)
Structure of apo human ferroportin in lipid nanodisc
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 6w4s
Q-score: 0.525
Billesbolle CB, Azumaya CM, Kretsch RC, Powers AS, Gonen S, Schneider S, Arvedson T, Dror RO, Cheng Y, Manglik A
Nature (2020) 586 pp. 807-811 [ Pubmed: 32814342 DOI: doi:10.1038/s41586-020-2668-z ]
- Ferroportin-fab45d8 complex (Complex)
- Fab45d8 heavy chain (23 kDa, Protein from Mus musculus)
- Fab45d8 light chain (24 kDa, Protein from Mus musculus)
- Fab45d8 (Complex from Mus musculus)
- Ferroportin (Complex from Homo sapiens)
- Solute carrier family 40 member 1 (66 kDa, Protein from Homo sapiens)
Structure of human ferroportin bound to hepcidin and cobalt in lipid nanodisc
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 6wbv
Q-score: 0.597
Billesbolle CB, Azumaya CM, Kretsch RC, Powers AS, Gonen S, Schneider S, Arvedson T, Dror RO, Cheng Y, Manglik A
Nature (2020) 586 pp. 807-811 [ Pubmed: 32814342 DOI: doi:10.1038/s41586-020-2668-z ]
- Cobalt (ii) ion (58 Da, Ligand)
- Ferroportin-fab45d8 complex (Complex)
- Fab45d8 heavy chain (23 kDa, Protein from Mus musculus)
- Fab45d8 light chain (24 kDa, Protein from Mus musculus)
- Fab45d8 (Complex from Mus musculus)
- Hepcidin (2 kDa, Protein from Homo sapiens)
- Solute carrier family 40 member 1 (66 kDa, Protein from Homo sapiens)
- Water (18 Da, Ligand)
- Ferroportin-hepcidin complex (Complex from Homo sapiens)
- (1s)-2-{[{[(2s)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(pentanoyloxy)methyl]ethyl octanoate (455 Da, Ligand)
Cryo-EM map of semi-attached mutant OCCM-DNA complex (ORC-Cdc6-Cdt1-Mcm2-7 with Mcm6 WHD truncation)
Resolution: 4.3 Å
EM Method: Single-particle
Fitted PDBs: 6wgc
Q-score: 0.301
Yuan Z, Schneider S, Dodd T, Riera A, Bai L, Yan C, Magdalou I, Ivanov I, Stillman B, Li H, Speck C
PNAS (2020) 117 pp. 17747-17756 [ Pubmed: 32669428 DOI: doi:10.1073/pnas.2006231117 ]
- Cell division control protein 6 (58 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 6 (50 kDa, Protein from Saccharomyces cerevisiae)
- Orc-cdc6-mcm3-mcm7-dsdna (Complex from Saccharomyces cerevisiae)
- Origin recognition complex subunit 5 (55 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 1 (104 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 3 (72 kDa, Protein from Saccharomyces cerevisiae)
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
- Dna replication licensing factor mcm3 (107 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm7 (95 kDa, Protein from Saccharomyces cerevisiae)
- Dna (41-mer) (12 kDa, DNA from Saccharomyces cerevisiae)
- Origin recognition complex subunit 2 (71 kDa, Protein from Saccharomyces cerevisiae)
- Dna (41-mer) (12 kDa, DNA from Saccharomyces cerevisiae)
- Origin recognition complex subunit 4 (60 kDa, Protein from Saccharomyces cerevisiae)
Cryo-EM map of mutant Mcm2-7 hexamer with Mcm6 WHD truncation
Resolution: 7.7 Å
EM Method: Single-particle
Fitted PDBs: 6wgf
Q-score: 0.064
Yuan Z, Schneider S, Dodd T, Riera A, Bai L, Yan C, Magdalou I, Ivanov I, Stillman B, Li H, Speck C
PNAS (2020) 117 pp. 17747-17756 [ Pubmed: 32669428 DOI: doi:10.1073/pnas.2006231117 ]
- Dna replication licensing factor mcm4 (105 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm2 (98 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm3 (107 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm7 (95 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm6 (113 kDa, Protein from Saccharomyces cerevisiae)
- Minichromosome maintenance protein 5 (86 kDa, Protein from Saccharomyces cerevisiae)
- Mcm2-7 (Complex from Saccharomyces cerevisiae)
Cryo-EM map of pre-insertion mutant OCCM-DNA complex (ORC-Cdc6-Cdt1-Mcm2-7 with Mcm6 WHD truncation)
Resolution: 8.1 Å
EM Method: Single-particle
Fitted PDBs: 6wgg
Q-score: 0.104
Yuan Z, Schneider S, Dodd T, Riera A, Bai L, Yan C, Magdalou I, Ivanov I, Stillman B, Li H, Speck C
PNAS (2020) 117 pp. 17747-17756 [ Pubmed: 32669428 DOI: doi:10.1073/pnas.2006231117 ]
- Cell division control protein 6 (58 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 4 (60 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 6 (50 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm4 (105 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 5 (55 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 1 (104 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 3 (72 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm2 (98 kDa, Protein from Saccharomyces cerevisiae)
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
- Dna replication licensing factor mcm3 (107 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm6 (113 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm7 (95 kDa, Protein from Saccharomyces cerevisiae)
- Dna (41-mer) (12 kDa, DNA from Saccharomyces cerevisiae)
- Orc-cdc6-mcm2-7-dsdna (Complex from Saccharomyces cerevisiae)
- Minichromosome maintenance protein 5 (86 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 2 (71 kDa, Protein from Saccharomyces cerevisiae)
- Dna (41-mer) (12 kDa, DNA from Saccharomyces cerevisiae)
- Cell division cycle protein cdt1 (68 kDa, Protein from Saccharomyces cerevisiae)